1Department of Agricultural Convergence Technology, Jeonbuk National University, Jeonju 54896, Republic of Korea
2Laboratory of Developmental Genetics, Department of Biomedical Sciences, Inha University College of Medicine, Incheon, 22212, Republic of Korea
3International Agricultural Cooperation and Development Center, Jeonbuk National University, Jeonju 54896, Republic of Korea
Correspondence to Ki-Duk Song, E-mail: kiduk.song@jbnu.ac.kr
Volume 9, Number 4, Pages 193-202, December 2025.
Journal of Animal Breeding and Genomics 2025, 9(4), 193-202. https://doi.org/10.12972/jabng.2025.9.4.2
Received on October 27, 2025, Revised on December 18, 2025, Accepted on December 18, 2025, Published on December 31, 2025.
Copyright © 2025 Korean Society of Animal Breeding and Genetics.
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0).
Hemopexin (HPX) is a plasma glycoprotein with a high affinity for heme, acting as a key antioxidant defense molecule against heme-induced oxidative stress. However, limited information is available on chicken HPX, particularly regarding genetic polymorphisms that may influence its biochemical properties. In this study, conserved motifs and post-translational modification sites were analyzed via Clustal Omega, NetNGlyc-1.0, and NetPhos-3.1. Along with that, non-synonymous single-nucleotide polymorphisms (nsSNPs), which were indicated as deleterious within the chicken HPX gene, were identified from the Ensembl genome database. The threedimensional structure of wild-type and mutant HPX proteins was modeled using AlphaFold3, and their interactions with the HEME ligand were evaluated through HADDOCK 2.4 and PRODIGY binding affinity prediction. The results show that high conservation of histidine residues is essential for HEME binding. Four N-glycosylation and twenty-four phosphorylation sites were predicted, suggesting regulatory complexity in HPX function. Among the identified nsSNPs, mutations such as L414T and A349V exhibited enhanced predicted binding affinities (ΔG = -13.35 and -13.33 kcal/mol, respectively) compared to the wild type (ΔG = -13.17 kcal/ mol). Therefore, for the variants in this study, which are located in the C-terminal β-propeller domain, some may not affect heme binding, while others may enhance heme binding. In conclusion, this study provides interesting insight for future research on HPX polymorphisms as potential biomarkers for oxidative stress resistance and selection markers in poultry production.
Chicken, Hemopexin, HEME binding, Molecular docking, Oxidative stress, SNPs
Hemoglobin circulating in the blood performs various vital functions in the body. However, if released into the bloodstream through hemolysis of red blood cells, it can cause potential toxicity. In particular, heme released from hemoglobin can be incorporated into cell membrane lipids, generating hydroxyl radicals, a powerful reactive oxygen species, which is a major cause of oxidative stress and endothelial dysfunction (Balla et al., 2005). These toxic substances can cause serious damage to cells and tissues. Various defense systems exist within the body to effectively remove the toxicity of hemoglobin-derived heme, and hemopexin (HPX) is a plasma protein that plays a key role among them (Dutra & Bozza, 2014; Vinchi & Tolosano, 2013).
HPX has the highest binding affinity for heme among plasma proteins, rapidly and efficiently capturing free heme (Smith & McCulloh, 2015). HPX, primarily synthesized in the liver and secreted into the blood, has been shown to increase its synthesis following inflammatory responses. The HPX-HEME complex binds to the HPX receptor (CD91) present on the surface of specific cells and is taken up into the cell. Heme is then degraded by heme oxygenase-1 to form substances such as biliverdin, which induce antioxidant activity. Therefore, the primary role of HPX is known to lie in its antioxidant action and cytoprotective function against heme overload following intravascular hemolysis, rather than in the regulation of iron metabolism (Tolosano et al., 2010; Tolosano & Fiorella, 2002). This constitutes an important mechanism for protecting the body from oxidative stress.
While research on the function of HPX is actively underway in various species, including mammals, little is known about the molecular biology and function of the HPX gene and protein in poultry, particularly chickens (Gallus gallus). Chickens occupy a crucial role in the poultry industry, and exposure to various stressors (e.g., heat stress, disease, environmental changes, etc.) can lead to hemolysis, leaving them vulnerable to oxidative stress induced by heme metabolites.
The HPX was suggested to play a role in protecting against oxidative stress and early mineralization during eggshell formation, causing its overexpression in the white isthmus, which was detected in proteomic analyses (Du et al., 2015). Riva et al.\(2024) suggested a new biomarker of the acute phase response (APR) in hens exposed to E. coli lipopolysaccharide (LPS) endotoxin is hemopexin. However, studies on HPX in poultry are still very limited. Through sequence analysis, they confirmed that the chicken HPX gene exhibits high homology with mammalian HPX, particularly the conservation of histidine residues, known to play a crucial role in heme binding (Dooley et al., 2010). These results suggest that chicken HPX may also perform biological functions similar to those of mammalian proteins.
Previous studies have identified chicken HPX as a developmentally regulated and acute-phase response protein. Grieninger et al.\(1986) demonstrated that HPX is produced within 12 hours of culture in embryonic chicken hepatocytes without hormones or other supplements, and its production increases rapidly, suggesting an important role in early development. Furthermore, Adler et al.\(2001) observed that when chickens were injected with antigens or lipopolysaccharide (LPS), which induce an acute phase response, plasma HPX and alpha-1-acid glycoprotein (AGP) levels significantly increased, as did HPX mRNA expression in the liver. These results strongly suggest that chicken plasma HPX or AGP levels can be used as indicators of the systemic response to local and systemic inflammations in the chicken.
Furthermore, as part of genetic diversity research, it is crucial to identify single-nucleotide polymorphisms (SNPs) and elucidate their functional impact. SNPs within genes can directly affect protein structure and function by altering amino acid sequences, or they can alter gene expression levels by altering transcriptional regulatory regions. In livestock such as chickens, SNP research is essential for identifying genetic markers associated with specific traits (e.g., productivity, disease resistance).
Considering these previous studies, to clearly elucidate the functional role of chicken HPX, in-depth basic research is urgently needed, including functional regulation by genetic mutations (SNPs) and the transcriptional regulatory mechanisms that control gene expression. Therefore, this study aims to assess the molecular biological function of genetic polymorphisms of the chicken HPX gene, based on bioinformatics predictions and physiological observations from previous studies. Ultimately, this research is expected to contribute to the development of new genetic breeding markers or disease control strategies to improve chicken health and productivity.
The hemopexin amino acid sequences of chicken (NP_001383363.1) and several species, including human (NP_000604.1), quail (XP_015708049.1), horse (XP_005612174.1), sheep (NP_001029784.1), rat (NP_059067.2), and mouse (NP_059067.2), were obtained from NCBI (https://www.ncbi.nlm.nih.gov/nucleotide/) in FASTA format. The chicken HPX nucleotide sequences (accession number: NP_001383363.1) were obtained and used to identify the SNPs. Chicken HPX SNPs information was scientifically retrieved from the Ensembl genome database (https://www.ensembl.org/index.html?redirect=no) with the gene identifier (ENSGALT00010003685.1) of the HPX gene in the Gallus gallus reference genome. All genetic variants associated with the HPX locus were found. Only missense variations were sorted to examine their impact on HEME binding.
Clustal Omega Website (https://www.ebi.ac.uk/jdispatcher/msa/clustalo) was used to perform alignment and neighbor-joining (NJ) phylogenetic tree of the HPX protein sequence between species (Madeira et al., 2024).
The Prediction of N-glycosylation and Phosphorylation sites were conducted using DTU Health Tech bioinformatics tools. NetNGlyc-1.0 (https://services.healthtech.dtu.dk/services/NetNGlyc-1.0/) was used to predict N-linked glycosation sites in the HPX wild-type protein. NetnGlyc uses artificial neural networks to examine the sequence context of Asn-Xaa-Ser/Thr sequons where Xaa is not Proline whether they are glycosylated and non-glycosylated asparagines (Gupta & Brunak, 2002). Analysis uses 0.5 as the default thresholds and the predicted value that revolve 0.5 are used to assign higher confidence levels for positive and negative sites.
Together, NetPhos-3.1 (https://services.healthtech.dtu.dk/services/NetPhos-3.1/), which uses neural networks to predict the serine, threonine, or tyrosine phosphorylation sites in both generic and kinase-specific in eukaryotic proteins (Blom et al., 2004). The prediction score ranged from 0.000 to 1.000, and the scores which above 0.500 indicate the potential positive predictions. Only positive predictions that have string YES were chosen. “unsp” indicated for residue site that predicted to be phosphorylated but couldn’t assign for any specific prediction.
AlphaFold3 Server (https://alphafoldserver.com/) was used to predict the three-dimensional structure of the protein〝ligand interaction model (Abramson et al., 2024). The model of each mutated HPX protein with HEME ligand was predicted in turn. The best model was chosen for further analysis based on the structural quality metrics which consist of the predicted Local Distance Difference Test (pLDDT) scores.
Molecular docking was performed to investigate the interactions between chicken SNP-contained HPX protein and HEME ligand. The molecular docking simulations were performed using HADDOCK 2.4 server (https://rascar.science.uu.nl/haddock2.4/). HADDOCK promotes the docking process and supports a variety of test data, including mutation data based on an integrated modeling approach (Honorato et al., 2024). The default docking parameters were used while the active residues involved in the interaction were manually determined. The active residue that defined for the binding site in HPX protein was assigned to conserved residues included PRO84, HIS87, HIS162, ARG209, TYR211, ARG220, TYR232, HIS248, HIS249.
PROtein binDIng enerGY prediction (PRODIGY, https://rascar.science.uu.nl/prodigy/) was employed to predict the binding affinity of the HPX-HEME docking complexes. PRODIGY is an advanced computational tool that predicts the binding strength of a given protein-protein complex based on the structural properties of its complex (Xue et al., 2016). Top-ranked HADDOCK complexes in PDB format were directly submitted to PRODIGY.
The liver is the primary source of chicken HPX synthesis, which is a vital HEME-binding glycoprotein in Gallus gallus, involved in immunological defense, detoxification, and the regulation of oxidative stress (Liebert, 2002). Not only acting as a major in heme transportation in plasma, which helps to prevent HEME-mediated oxidative stress, HPX has also been investigated as a potential biomarker for the Acute Phase Response (APR) in broilers using an E. coli LPS experimental model (Riva et al., 2024). The similarity of HPX amino acid sequences between chicken and those from other species was analyzed by bioinformatics tools, which allow identification of conserved and variable regions by creating the protein sequence alignment (Fig. 1).
The alignment of multiple HPX amino acid sequences from chicken and others revealed both conserved and variable regions (Fig. 1). The alignment result indicates that the chicken HPX protein shared the highest similarity with quail. Although all HPX mammalian sequences in this study exhibited a high degree of conservation among themselves, a greater divergence was observed in avian sequences (chicken and quail). Furthermore, amino acid substitutions and a number of gaps were observed between species, suggesting that the HPX protein underwent evolutionary adaptation while retaining its conserved functional motifs.
These results are also shown through the phylogenetic tree that HPX proteins from mammalian and poultry were clustered into two separate branches (Fig. 2). In the cluster of poultry, the amino acid sequences of chicken and quail were clustered together, showing the close relationship between them. Meanwhile, mammalian clusters include the remaining species and the HPX protein of rat and mouse clustered into a sub-branch.
The phylogenetic tree was established by multiple amino acid sequence alignment of HPX proteins from various species. The graph exhibits the evolutionary distance and relationship level among species.
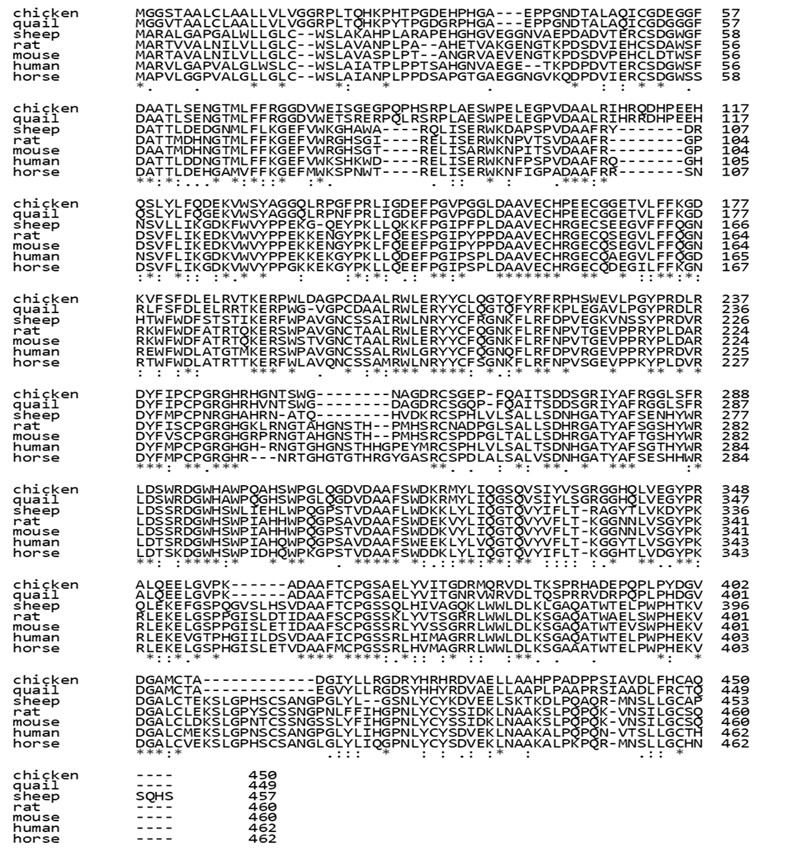
Figure 1. Investigating Amino Acid Sequence Similarity using Clustal Omega. HPX proteins were accessed for the similarity to investigate the modification in sequence between species.
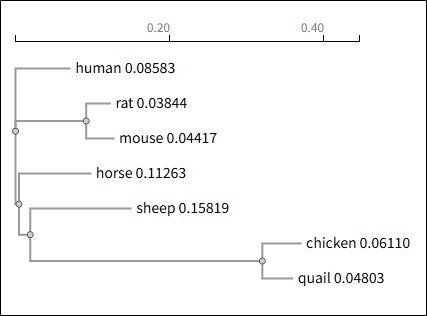

Figure 2. Phylogenetic tree of HPX protein among species.
Protein function is considerably modified by the covalent process known as protein post-translational modification (PTM), which happens in proteins during or after translation and involves the addition or removal of one or several functional groups. The result of N-glycosation prediction in HPX protein was presented in Table 1. There are 4 positions that SEQUON ASN-XAA-SER/THR, where XAA could be any amino acid except Proline (P), have been predicted as N-glycosylated sites. All of them had a threshold result greater than 0.5, and the jury agreement, which indicates how many of the nine networks support the prediction, was also satisfied. Notably, positions 65-NGTM and 257-NAGD exhibited the highest confidence levels for positive predicted sites. Missense mutations which is even near the N-glycosylation sites could prevent glycosylation or disrupt the structurally critical domains (Wei et al., 2017). This may directly influence the protein’s stability and biological function in plasma.
Along with that, HPX chicken protein is also predicted for phosphorylation sites by NetPhos-3.1 (Table 2). Twenty-four phosphorylation sites were predicted in the wild type of HPX sequence, means that these amino acid residues could be phosphorylated under certain biological circumstances. The predictions were tested for 17 specific kinases; however, the majority of the predictions are unknown. Means that, although it is predicted that these sites will be phosphorylated, the prediction model was unable to assign them to a particular family of protein kinases.
Table 1. Prediction of N-glycosylation sites in chicken HPX protein
| Sequence | Position | Potential | Jury agreement | N-Glyc result | |
|---|---|---|---|---|---|
| Chicken HPX | 43 | NDTA | 0.6208 | 7/9 | +* |
| 65 | NGTM | 0.7294 | 9/9 | ++ | |
| 252 | NTSW | 0.5273 | 5/9 | + | |
| 257 | NAGD | 0.6738 | 9/9 | ++ | |
*+: positive
Table 2. Prediction of phosphorylation sites in chicken HPX protein
| No. | Sequence | Position | Residue | Context | Score | Kinase |
|---|---|---|---|---|---|---|
| 1 | chicken | 4 | S | -MGGSTAAL | 0.677 | PKA1 |
| 2 | chicken | 23 | T | GRPLTQHKP | 0.868 | unsp |
| 3 | chicken | 29 | T | HKPHTPGDE | 0.828 | unsp |
| 4 | chicken | 80 | S | VWEISGEGP | 0.571 | unsp |
| 5 | chicken | 94 | S | PLAESWPEL | 0.993 | unsp |
| 6 | chicken | 170 | T | CGGETVLFF | 0.631 | PKC2 |
| 7 | chicken | 181 | S | DKVFSFDLE | 0.552 | CKII3 |
| 8 | chicken | 189 | T | ELRVTKERP | 0.501 | unsp |
| 9 | chicken | 225 | S | FRPHSWEVL | 0.997 | unsp |
| 10 | chicken | 254 | S | HGNTSWGNA | 0.981 | unsp |
| 11 | chicken | 263 | S | GDRCSGEPF | 0.857 | unsp |
| 12 | chicken | 272 | S | QAITSDDSG | 0.691 | unsp |
| 13 | chicken | 275 | S | TSDDSGRIY | 0.971 | unsp |
| 14 | chicken | 286 | S | RGGLSFRLD | 0.749 | PKC2 |
| 15 | chicken | 291 | S | FRLDSWRDG | 0.997 | unsp |
| 16 | chicken | 317 | S | DAAFSWDKR | 0.716 | PKC2 |
| 17 | chicken | 328 | S | LIQGSQVSI | 0.608 | DNAPK4 |
| 18 | chicken | 333 | Y | QVSIYVSGR | 0.502 | unsp |
| 19 | chicken | 335 | S | SIYVSGRGG | 0.764 | unsp |
| 20 | chicken | 372 | Y | SAELYVITG | 0.986 | unsp |
| 21 | chicken | 375 | T | LYVITGDRM | 0.515 | CDK15 |
| 22 | chicken | 385 | T | RVDLTKSPR | 0.678 | PKC2 |
| 23 | chicken | 387 | S | DLTKSPRHA | 0.986 | unsp |
| 24 | chicken | 420 | Y | RGDRYHRHR | 0.846 | unsp |
1PKA: Protein Kinase A, 2PKC: Protein Kinase C, 3CKII: Casein Kinase II,
4DNAPK: DNA Dependent Kinase, 5CDK1: Cyclin Dependent Kinase 1
Using the Ensembl platform, which provides publicly available genomic annotation data, SNP information for the chicken HPX gene in the Gallus gallus reference genome was identified (Dyer et al., 2025). The detected variants included both synonymous and missense substitutions. Missense SNPs were selected for downstream analyses because of their potential to alter protein structure and affect HEME-binding function. The identified missense variants are summarized in Table 3. For each Variant ID, the chromosome coordinates are listed along with the annotation of the polymorphism on the allele that results in the amino acid change. Additionally, based on the SIFT results with a threshold of 0.05, variants with results below the threshold are predicted to be deleterious〞possibly damaging. SIFT is used as a tool for predicting the impact of amino acid substitutions on protein function based on sequence homology and physical properties, applied to both naturally occurring polymorphisms and laboratory-induced missense mutations. On the other hand, MutPred2 provided complementary values by predicting the pathogenic potential of each variant with a threshold of 0.5. In the case of variant at residue 210, D201L and D201V (rs738341821), predicted results show that the mutations could disrupt the HPX function protein, caused by both low SIFT scores (0.02) and high MutPred2 scores (>0.8). Similarly, variant rs732484061 at residue 414 also has high MutPred2 scores (0.729〝0.830), predicted to be damaging. Conversely, some mutations, such as rs313782174, rs737170979, have MutPred2 scores that do not exceed 0.5, suggesting that they might be neutral in terms of protein function.
Table 3. Non-synonymous SNPs and functionality of the chicken HPX variants
| Residue | Variant ID | Chr: bp | ConseqType | Alleles | Mutation | Codons | SIFT | MutPred2 score |
|---|---|---|---|---|---|---|---|---|
| 132 | rs313782174 | 1:196135806 | missense variant | T/G | A132E | GAG | 0.04 | 0.321 |
| 201 | rs738341821 | 1:196135479 | missense variant | A/C | D201L | TTG | 0.02 | 0.857 |
| 201 | rs738341821 | 1:196135479 | missense variant | A/C | D201V | GTG | 0.824 | |
| 343 | rs740322613 | 1:196134103 | missense variant | T/C | V343R | AGA | 0.01 | 0.469 |
| 343 | rs740322613 | 1:196134103 | missense variant | T/C | V343G | GGA | 0.552 | |
| 349 | rs737170979 | 1:196134084 | missense variant | A/C | A349V | GTA | 0 | 0.109 |
| 349 | rs737170979 | 1:196134084 | missense variant | A/C | A349G | GGA | 0.107 | |
| 414 | rs732484061 | 1:196133811 | missense variant | T/G | L414T | ACA | 0.01 | 0.729 |
| 414 | rs732484061 | 1:196133811 | missense variant | T/G | L414P | CCA | 0.830 |
The mutation, non-synonymous SNPs that are located in highly conserved regions are more dangerous than found in non-conserved areas causing modification on structure and leading to disease of missing function. The study focused on investigating protein-ligand interactions in order to identify connections between wild-type and mutated variations to HEME ligand. Each protein-ligand model was predicted using AlphaFold 3 and the highly confidence model was chosen for conducting docking analysis. The molecular docking analysis was completed using the HADDOCK 2.4 server to evaluate the interaction of HPX-HEME complexes. The active residue that defined for the binding site in HPX protein was assigned to 9 conserved residues consists of 4 residues HIS87, HIS162, HIS248, HIS249 which was determined by similarity alignment compared to human HPX (P02790) and HIS also being reported as important residue that relevant to the heme binding (Lechuga et al., 2022). HADDOCK created around 200 models for each HPX-HEME interaction and isolated them into cluster that each model in the same cluster will exhibit similar interactions. In this study, the protein-ligand model that attains the highest score, best fit docking position, smaller RMSD and larger cluster size were selected and continued evaluated based on their binding affinity (利G) through the PRODIGY server. Protein-ligand docking and 利G assessment were performed one by one and the results were presented in the Table 4. Docking model of HPX-HEME was shown on Figure 3.
Using HADDOCK 2.4 and PRODIGY, the docking and binding affinity analyses showed that the chicken HPX variants differed significantly in their protein-heme interactions. The docking structure of the HPX-HEME complex shows the similar binding area between two similar four-bladed 乒-propeller domains that is surrounded by an interdomain linker (Paoli et al., 1999). The overall quality of each docking is generally evaluated through the HADDOCK score, where the more negative the score, the stronger binding. Table 4 shows that the docking result from several SNPs variants are stronger than the wild type (-87.0). Variants docking result of L414T (-106.9), A349V (-103.9), A349G (-97.9), V343G (-97.8) suggest that mutations around residues 343〝414 which located in C-terminal domain enhance heme affinity. The RMSD values range from 0.2 to 0.4Å with the standard deviation up to 0.3Å indicated that all models are structurally consistent and dependable.
The amino acid alteration that comes from nsSNPs could effect to the protein function and might directly impact to the protein-ligand interaction caused the change of the binding pocket environment or the protein’s surface charge distribution (Dakal et al., 2017; Sivakumar & Subbiah, 2024). Both the wild-type and mutant HPX proteins were able to form stable complexes with heme in the present study, but the binding strengths were different, according to docking and binding free energy. When compared to the wild type (-13.17 kcal/mol), the projected binding free energy (汁G) values varied from -13.35 to -11.24 kcal/mol, suggesting that certain mutations either marginally increased or decreased HEME affinity. Some variants such as D201V (-11.67 kcal/mol) and L414P (-11.24 kcal/mol) showed decreased affinity, means destabilization of the HPX-heme interaction. Besides that, A132E (-13.24 kcal/mol), A349V (-13.33 kcal/mol), and L414T (-13.35 kcal/mol) displayed the predicted binding energy 汁G values slightly increased when compared to the wild-type protein. It can be seen that Van der Waals interaction of A349V and L414T were -28,5 and -35,5 kcal/mol which helps heme more stabilizing in the binding area. In constrat with D201V which is complicated in binding with heme ligand caused the lowest Van der Waals value (-7,8 kcal/mol) and the huge electrostatic energy (-352,1 kcal/mol).
Overall, these results validate that the stabilization of HPX-HEME complex being affected due to SNP-induced amino acid substitutions. Although nsSNPs A132E, A349V and L414T show strong HEME-binding performance, other vaariants such as D201V and L414P may reduce the binding efficiency. These results demonstrate how sensitive the HPX-HEME interaction is to deleterious nsSNP, which may ultimately affect HPX’s physiological function in regulating heme transport in chicken.
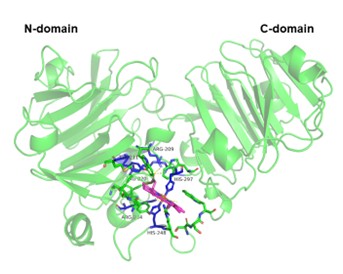
Figure 3. HPX-HEME docking model. 乒-propeller domain (green) from left to right are N-terminal domain and C-terminal domain of HPX protein, HEME ligand (magenta), critical residue (blue). The figure was rendered using PyMOL (Schrödinger, 2015).
Table 4. Result of docking and protein modeling
| Docking parameter | Wild type | Binding of HPX variants to HEME | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| A132E | D201L | D201V | V343R | V343G | A349V | A349G | L414T | L414P | ||
| HADDOCK score | -87.0+/-15.8 | -94.4+/-0.4 | -91.2+/-10.9 | -81.0+/-2.2 | -84.3+/-11.2 | -97.8+/-6.1 | -103.9+/-25.5 | -97.9+/-7.6 | -106.9+/-0.7 | -83.1+/-8.0 |
| 汁G prediction (Kcal/mol) | -13.17 | -13.24 | -12.97 | -11.67 | -12.43 | -12.63 | -13.33 | -12.91 | -13.35 | -11.24 |
| Cluster size | 93 | 145 | 100 | 101 | 92 | 176 | 178 | 124 | 162 | 158 |
| RMSD from the overall lowest-energy structure | 0.3+/-0.2 | 0.2+/-0.1 | 0.2+/-0.2 | 0.4+/-0.3 | 0.2+/-0.1 | 0.4+/-0.3 | 0.3+/-0.2 | 0.2+/-0.1 | 0.2+/-0.1 | 0.3+/-0.1 |
| Van der Waals energy | -22.4+/-11.2 | -25.7+/-9.3 | -25.6+/-11.5 | -7.8+/-7.6 | -26.6+/-8.4 | -36.5+/-2.7 | -28.5+/-10.4 | -32.5+/-5.4 | -35.5+/-0.7 | -24.4+/-7.9 |
| Electrostatic energy | -283.9+/-19.6 | -271.8+/-31.1 | -254.9+/-80.4 | -352.1+/-50.6 | -200.6+/-18.6 | -223.1+/-21.0 | -292.6+/-89.6 | -255.9+/-13.1 | -282.8+/-6.8 | -193.6+/-68.0 |
| Desolvation energy | -10.5+/-6.0 | -15.8+/-7.6 | -16.9+/-8.5 | -6.3+/-1.7 | -19.1+/-1.6 | -22.7+/-1.2 | -19.9+/-1.3 | -19.8 +/- 1.9 | -21.6+/-0.6 | -21.6+/-3.9 |
| Restraints violation energy | 26.3+/-44.9 | 14.8+/-24.7 | 22.8 +/- 21.5 | 35.1+/-26.0 | 15.9+/-22.7 | 59.4+/-7.5 | 29.7+/-27.9 | 55.5 +/- 37.3 | 67.0+/-23.1 | 16.8+/-26.7 |
| Buried Surface Area | 809.4+/-133.7 | 900.4+/-134.5 | 871.2+/-136.7 | 658.2+/-63.1 | 905.4+/-67.6 | 1049.4+/-51.4 | 957.0+/-43.7 | 994.9 +/- 59.5 | 1022.0+/-10.2 | 936.9+/-51.4 |
| Z-Score | -1.2 | -1.2 | -1.3 | -1.5 | -1.9 | -1.4 | -1.4 | -1.9 | -1.7 | -1 |
Although several missense SNPs were identified within the chicken HPX gene, structural modeling and docking analyses demonstrated that most variants do not negatively influence HEME-binding function. Notably, mutations such as L414T and A349V enhanced binding affinity and surface interactions with HEME, suggesting potential functional advantages in oxidative stress regulation. These findings indicate that specific HPX variants may contribute to adaptive mechanisms in avian physiology and could be developed as functional genetic markers for oxidative stress tolerance and immune resilience in chicken breeding programs.
This research was supported by the Regional Innovation System & Education(RISE) program through the Jeonbuk RISE Center, funded by the Ministry of Education(MOE) and the Jeonbuk State, Republic of Korea.(2025-RISE-13-JBU).
No potential conflict of interest relevant to this article is reported.
Abramson, J., Adler, J., Dunger, J., Evans, R., Green, T., Pritzel, A., Ronneberger, O., Willmore, L., Ballard, A. J., Bambrick, J., Bodenstein, S. W., Evans, D. A., Hung, C. C., O*Neill, M., Reiman, D., Tunyasuvunakool, K., Wu, Z., Žemgulytė, A., Arvaniti, E. Jumper, J. M. 2024. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature, 630(8016), 493-500.
[DOI][PubMed][PMC]
Abramson J, Adler J, Dunger J, Evans R, Green T, Pritzel A, Ronneberger O, Willmore L, Ballard AJ, Bambrick J, et al.\2024. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature. 630(8016):493-500.
[DOI][PubMed][PMC]
Adler KL, Peng PH, Peng RK, Klasing KC. 2001. The kinetics of hemopexin and 丟1-acid glycoprotein levels induced by injection of inflammatory agents in chickens. Avian Dis. 45(2):289-296.
[DOI][PubMed]
Balla J, Vercellotti GM, Jeney V, Yachie A, Varga Z, Eaton JW, Balla G. 2005. Heme, heme oxygenase and ferritin in vascular endothelial cell injury. Mol Nutr Food Res. 49(11):1030-1043.
[DOI][PubMed]
Blom N, Sicheritz-Pontén T, Gupta R, Gammeltoft S, Brunak S. 2004. Prediction of post-translational glycosylation and phosphorylation of proteins from amino acid sequence. Proteomics. 4(6):1633-1649.
[DOI][PubMed]
Dakal TC, Kala D, Dhiman G, Yadav V, Krokhotin A, Dokholyan NV. 2017. Predicting functional consequences of non-synonymous single nucleotide polymorphisms in IL8 gene. Sci Rep.\7(1):1-18.
[DOI][PubMed][PMC]
Dooley H, Buckingham EB, Criscitiello MF, Flajnik MF. 2010. Emergence of the acute-phase protein hemopexin in jawed vertebrates. Mol Immunol. 48(1-3):147-152.
[DOI][PubMed][PMC]
Du J, Hincke MT, Rose-Martel M, Hennequet-Antier C, Brionne A, Cogburn LA, Nys Y, Gautron J. 2015. Identifying specific proteins involved in eggshell membrane formation using gene expression analysis and bioinformatics. BMC Genomics. 16(1):1-13.
[DOI][PubMed][PMC]
Dutra FF, Bozza MT. 2014. Heme on innate immunity and inflammation. Front Pharmacol. 5(May):1-20.
[DOI][PubMed][PMC]
Dyer SC, Austine-Orimoloye O, Azov AG, Barba M, Barnes I, Barrera-Enriquez VP, Becker A, Bennett R, Beracochea M, Berry A, et al. 2025. Ensembl 2025. Nucleic Acids Res. 53(D1):D948-D957.
[DOI][PubMed][PMC]
Grieninger G, Liang TJ, Beuving G, Goldfarb V, Metcalfe SA, Müller-Eberhard U. 1986. Hemopexin is a developmentally regulated, acute-phase plasma protein in the chicken. J Biol Chem. 261(33):15719-15724.
[DOI][PubMed]
Gupta R, Brunak S. 2002. Prediction of glycosylation across the human proteome and the correlation to protein function. Pac Symp Biocomput. 7:310-322.
[DOI][PubMed]
Honorato RV, Trellet ME, Jiménez-García B, Schaarschmidt JJ, Giulini M, Reys V, Koukos PI, Rodrigues JPGLM, Karaca E, van Zundert GCP, et al. 2024. The HADDOCK2.4 web server for integrative modeling of biomolecular complexes. Nat Protoc. 19(Nov):1-14.
[DOI][PubMed]
Liebert MA. 2002. Hemopexin: structure, function, and regulation. Antioxid Redox Signal. 21(4):297-306.
[DOI][PubMed]
Madeira F, Madhusoodanan N, Lee J, Eusebi A, Niewielska A, Tivey ARN, Lopez R, Butcher S. 2024. The EMBL-EBI Job Dispatcher sequence analysis tools framework in 2024. Nucleic Acids Res. 52(W1):W521-W525.
[DOI][PubMed][PMC]
Paoli M, Anderson BF, Baker HM, Morgan WT, Smith A, Baker EN. 1999. Crystal structure of hemopexin reveals a novel high-affinity heme site formed between two 乒-propeller domains. Nat Struct Biol. 6(10):926-931.
[DOI][PubMed]
Riva F, Eckersall PD, Chadwick CC, Chadwick LC, McKeegan DEF, Peinado-Izaguerri J, Bruggeman G, Hermans D, McLaughlin M, Bain M. 2024. Identification of novel biomarkers of acute phase response in chickens challenged with Escherichia coli lipopolysaccharide endotoxin. BMC Vet Res. 20(1):1-8.
[DOI][PubMed][PMC]
Schrödinger LLC. 2015. The PyMOL Molecular Graphics System, Version 1.8.
Sivakumar K, Subbiah U. 2024. Computational analysis of non-synonymous SNPs in the human LCN2 gene. Egypt J Med Hum Genet. 25(1):1-10.
[DOI]
Smith A, McCulloh RJ. 2015. Hemopexin and haptoglobin: Allies against heme toxicity from hemoglobin, not contenders. Front Physiol. 6:187.
[DOI]
Tolosano E, Fiorella A. 2002. Hemopexin: structure, function, and regulation. Antioxid Redox Signal. 21(4):297-306.
[DOI][PubMed]
Tolosano E, Fagoonee S, Morello N, Vinchi F, Fiorito V. 2010. Heme scavenging and the other facets of hemopexin. Antioxid Redox Signal. 12(2):305-320.
[DOI][PubMed]
Vinchi F, Tolosano E. 2013. Therapeutic approaches to limit hemolysis-driven endothelial dysfunction: Scavenging free heme to preserve vasculature homeostasis. Oxid Med Cell Longev. 2013:396527.
[DOI][PubMed][PMC]
Wei W, Zheng C, Zhu M, Zhu X, Yang R, Misra S, Zhang B. 2017. Missense mutations near the N-glycosylation site of the A2 domain lead to various intracellular trafficking defects in coagulation factor VIII. Sci Rep. 7:45033.
[DOI][PubMed][PMC]
Xue LC, Rodrigues JP, Kastritis PL, Bonvin AM, Vangone A. 2016. PRODIGY: a web server for predicting binding affinity of protein-protein complexes. Bioinformatics. 32(23):3676-3678.
[DOI][PubMed]